Quality in ART – Innovation or back to basics?
The ESHRE 2020 Scientific Programme presented a mixture of back to basics approaches and innovation. For this edition of our newsletter it has been a great pleasure to summarize some of the key insights our scientific team identified, including predictions on the future of IVF!
Back to Basics
Big data, better trials
The importance of being able to effectively collect and analyze our data featured prominently. Case studies, observational data, randomized controlled trials (RCTs), analysis of national and international databases – all are valid. But as S Nelson (O-001) highlighted, we need to think big and be aware of the numbers needed to prove statistically significant benefits.
M van Wely (O-219) later expanded on Nelson’s call for appropriate guidelines, including enhanced recruitment, better quality (with better adherence and monitoring), and effective follow-up in ART studies, discussing when RCTs are needed and how they should be implemented. They concluded it is OK to do an RCT on a patient population in which the intervention is expected to work and a larger effect size means fewer patients are needed. Once benefit is established, the intervention might be tested in other populations or subgroups. The difference and discrepancies between RCTs and Cochrane reviews were highlighted – poor data or pooled data from studies with methodological issues can give exaggerated differences.
Embryology & Andrology
In a pre-congress course, R Sturmey (PC-17) gave a comprehensive review of embryo metabolism and the requirements of the constituents of culture media. The long-held belief that increased glucose consumption during blastocyst activation is to satisfy energy demands is incorrect – glucose is essential for compaction but does not contribute to the Krebs cycle. Rather, oxygen consumption rises to meet the demand for ATP during development.
Meanwhile, the epigenetic landscape is influenced by maternal metabolic physiology and biochemical modification at the cleavage stage, which changes the epigenome and transcriptome in the blastocyst. The central role of pyruvate and glucose under differing oxygen demands during development was highlighted, which is of particular importance given not all culture media provide the same concentrations of these essential nutrients.
S Brouillet’s (O-288) presentation in the Scientific Programme on dynamic oxygen levels during human embryo culture underlined Sturmey’s point. 5% O2 was used up to D3, then reduced to 2%, resulting in significantly improved usable blastocyst rate and cumulative live birth rates.
The constant debate concerning ICSI versus IVF continued with QV Dang (O-167) examining whether ICSI results in a higher ongoing pregnancy rate compared to conventional IVF in couples with non-male factor infertility.
The study showed ICSI did not result in a higher fertilization rate, lower total fertilization failure or higher ongoing pregnancy rate compared to conventional IVF, questioning the value of standard use of ICSI in non-male infertility in routine assisted reproduction.
N De Munck (O-166) examined differences in the morphokinetic behavior between IVF and ICSI embryos in PGT-A cycles. Preimplantation development is marginally yet significantly different between embryos generated by IVF and ICSI. When standardizing for the time of pronuclear fading, the differences in early cleavage disappear between IVF and ICSI, while blastulation and blastocyst expansion occur earlier for IVF embryos.The study showed conventional IVF results in the same number of blastocysts for biopsy with similar developmental kinetics, thereby reinforcing the use of conventional IVF in this patient population. Despite this, C Wyns (O-044) presented preliminary data from European registers by the ESHRE EIM Consortium, demonstrating an ongoing trend towards increased ICSI usage.
Fresh versus frozen embryo transfer
Several studies employed big data to try and answer the question of whether fresh or frozen is best. C Wyns (O044) and GD Adamson (O-0157) confirmed European and global trends towards freeze-all cycles and frozen embryo transfers (FETs).
K Westvik-Johari (O-029) presented data from a Nordic sibling study comparing fresh and frozen IVF cycles to spontaneous conceptions and demonstrated that fresh ETs resulted in the lowest birth weight and frozen ETs resulted in the highest, but both ART groups were at higher risk of pre-term birth.
A Terho (O-031) also presented data from a Nordic study, again showing higher birth weights in FET babies, beginning at 33 weeks for boys and 34 weeks for girls, with the difference in the number of babies being large for gestational age becoming significant at 36 weeks.
L Vuong (O-032) showed data suggesting that children born from freeze-only versus fresh embryo transfer demonstrated better problem solving and motor skills at two years of age.
Preliminary data from the UK E-Freeze RCT was presented by A Maheshwari (O-061). From this, a reduction in ovarian hyperstimulation syndrome (OHSS) and an improved live birth rate in hyper-responders has been observed. Overall, increased time to pregnancy and a higher rate of large for gestational age babies was also observed alongside a lower rate of small for gestational age babies and preterm deliveries.
Elsewhere in the main programme N Aslih (O-172) demonstrated natural cycle FET could almost double the clinical pregnancy rate compared to artificial endometrial preparation, even where embryo quality was poor.
G Oron (O-255) indicated the clinical pregnancy rate of the first frozen-thawed transfer is higher if a clinical pregnancy was attained in the fresh transfer regardless of live birth. Identifying predictive factors for the success of frozen-thawed embryo transfers is important to maintain acceptable pregnancy rates while reducing multifetal pregnancies and can help physicians while counseling patients regarding the number of embryos to transfer, taking into consideration the outcome of the previous fresh cycle.
INNOVATIONS
Andrology
After years of seeming neglect, there was a focus on novel sperm tests and improving diagnostics to anticipate success or failure of treatment cycles and intervene as necessary. E Baldi (O-099) called into question the predictive value of semen analysis and asserted that reliance on ICSI had overtaken efforts to find better andrological tests.
There is the potential for capacitation as a possible marker, measurement of membrane potential, and assessment of hyperactivation. Additionally, CatSper has been linked to embryo quality and gene expression, along with measurement of chromatin protamination, and may be a good predictor of outcomes.
While advanced tests may prove useful, most remain difficult to perform, and simple, point-of-care tests are required.
Several presentations examined genetic tests and markers. D Lamb (O-100) examined the role of genetic testing on the evaluation of male infertility. As sperm count declines, the incidence of genetic abnormalities broadly increases. In addition to standard recommendations – karyotypes and Cystic Fibrosis mutation panels – teratozoospermia and globozoospermia are shown to be linked to various deletions, mutations, and polymorphisms.
Next generation sequencing allows easier analysis of panels of genes associated with male infertility, sperm dysfunctions, and defects of the genitourinary tract as well as emission/ejaculation disorders which could be important for genetic counseling and understanding/ predicting poorer treatment outcomes for some patients.
S Cheung (O-272) examined germline mutations in azoospermic men and related them to etiology and outcomes. D Tavares (O-297) demonstrated globozoospermia to be associated with specific gene mutations and used bioassays to assess the sperm’s capacity to support embryonic development and tailor treatment.
Related to this, P Chung (O-299) investigated screening the male gamete for the presence of phospholipase C zeta as a means of identifying the specific gamete responsible for ICSI complete fertilization failure and guide treatment in subsequent cycles.
Similar to Tavares’s findings, where a sperm-related oocyte activation deficiency was confirmed, calcium ionophore treatment improved outcomes, with offspring displaying normal development at three years of age. Chung also demonstrated that where testing did not point to a sperm defect, outcomes can be improved with tailored superovulation protocols.
I Arenaza (O-018) used proteomic analysis via mass spectrometry to identify normozoospermic men with unexplained infertility less likely to achieve implantation and pregnancy.
P Schlegel (O-052) also cited semen analysis as a poor measure of male fertility, with lifestyle factors, obesity, and abstinence period having a significant impact on sperm quality and ART outcomes, and DNA fragmentation being a better predictor.
C Depuydt (O-017) noted IUI sperm samples with human papillomavirus (HPV) had a higher DNA fragmentation index (DFI) and that clinical pregnancies were only observed with a DFI <26% and in the absence of HPV infection. Testing for HPV and sperm DFI could help differentiate between those patients that would be best treated by IUI or IVF/ICSI.
A Melnick (O-246) investigated a previous study from Xie et al, which mapped DNA fragmentation in sperm from different parts of the male genital tract, to determine which region of the epididymis yields sperm resulting in best embryo development. Fertilization, implantation rate, clinical pregnancy rate and live birth rate all followed the same pattern: cauda > corpus > caput which appears to reflect epididymal maturation. Also related to surgical sperm retrieval, V Vloeberghs (O-126) demonstrated the added value of enzymatic digestion in testicular sperm retrieval in non- obstructive azoospermia, increasing the percentage of patients with sperm found from 16.2% to 43.3%.
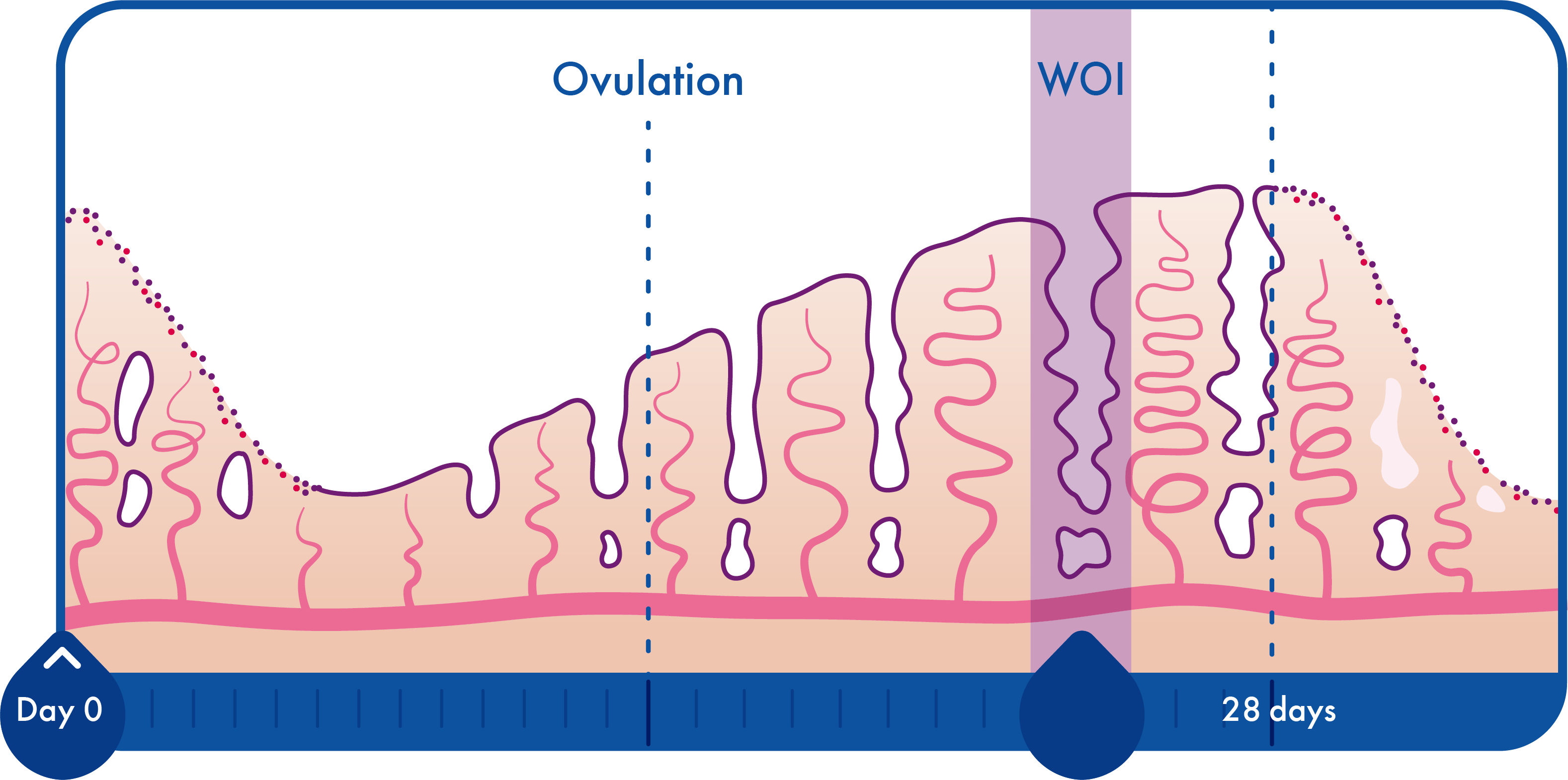
Endometrial Receptivity
Many studies demonstrated extended utility for molecular endometrial testing.
L Craciunas (O-202) showed that abnormal endometrial transcriptomic profiles build on the evidence to support nonchromosomal causes for higher- order miscarriages.
When compared to controls (two or three miscarriages), women who suffered four or more miscarriages had 19 differently expressed genes after adjustment for multiple comparisons. They were related to biological processes such as immunity, energy production, hormone secretion, adhesion, and cell proliferation.
When women who suffered a subsequent miscarriage were compared to women who had a live birth, following the endometrial biopsy, the number of significantly differentially expressed genes was 421, mapping to processes relevant to embryo implantation and early stages of pregnancy such as biosynthetic process, signal transduction, response to stress, immune system process, cell differentiation, catabolism, cell death, cell proliferation, homeostatic process, cytoskeleton organization, cell adhesion and reproduction.
From the same research group, J Chu (O-204) demonstrated abnormal endometrial metabolomic profiles may also build on the evidence to support nonchromosomal causes for higher-order miscarriages.
Again, there were distinct metabolomic profiles between the endometrial samples obtained from women with higher-order miscarriages, and changes were also identified in the metabolomic profiles of women who suffered a subsequent miscarriage compared to those who had a live birth following the endometrial biopsy.
Validation of the findings from these studies may aid the development of a predictive test for use in clinical practice.
P Ruane (O-032) described how receptive endometrial epithelium induces trophectoderm differentiation to invasive syncytiotrophoblast, which initiates invasive implantation. Further research and understanding may lead to embryo assessment for syncytiotrophoblast differentiation propensity pre-implantation or assessment of endometrial epithelium for propensity to induce syncytiotrophoblast differentiation.
Embryology
There are conflicting studies regarding the suitability of mtDNA as a biomarker of embryo implantation potential. M Galain (O-314) observed a significant difference in mtDNA in terms of ploidy and maternal age but not in terms of implantation rates, pregnancy rates, miscarriage rates or live birth rates, suggesting it does not appear to be a clinically useful biomarker.
J Mertens (O-138) found significant differences in the mitochondrial genome of children born after ART, which relates to a background of maternal infertility and birth weight. These results suggest a link between maternal infertility and mtDNA variant composition, which is transmitted to the offspring.
C Lin (O-064) determined that H3.3 Chaperone Hira complex (present in the oocyte) is essential for male pronucleus formation. In the loss-of-function mouse models abnormal fertilization occurred and formation of a single pronucleus (1PN).
In human 1PNs it was observed that Hira chaperone molecules failed to incorporate into male chromatin. This suggested possible development of “1PN rescue” using overexpression, or nuclear transfer approaches for patients with repeated 1PN abnormal fertilization.
K Ezoe (O-063) identified prolactin (PRL) as a potential therapeutic target and developed supplemented media. PRL is thought to influence preimplantation development and implantation. Human embryos do not appear to express the prolactin receptor (PRLR) before the morula stage – PRLR signaling stimulates blastocyst adhesion.
The supplementation of the embryo culture medium with PRL did not improve the rate of embryonic development to the blastocyst stage or their morphological grade but significantly increased blastocyst outgrowth. PRL treatment during embryo culture could be advantageous for improving pregnancy outcomes following blastocyst transfer.
Automation, AI and novel technology
In the pre-congress courses, D Sakkas (PC-16) reflected that PGT-A and morphokinetics do not tell us everything about embryo viability and pointed to a lab of the future where non-invasive techniques will eventually replace invasive techniques for assessing embryo viability, including genetic “normality”.
Fluorescence-lifetime imaging microscopy (FLIM) can provide information on the auto-fluorescence of NADH and FAD, important intermediaries in the electron transport chain, and reveals dramatic differences in FAD within the ICM and TE. Hyperspectral microscopy similarly can investigate auto-fluorescent metabolites.
Raman micro-spectroscopy, along with FLIM, has been applied to differentiate euploid and aneuploid embryos. In the future, these non-invasive platforms may be coupled with microfluidics for live-time imaging of individual embryo metabolomics, which could enable a fluid, dynamic system of embryo culture, analysis, and control of development.
A Adjuk (O-310) demonstrated that in addition to visualizing nuclei within morulae, optical coherence microscopy can allow assessment of chromatin conformation in germinal vesicles (GVs) and the number of cells in morulae, and that these correlate with developmental potential.
Following studies showing the ability of convolutional neural networks to identify morphologically normal sperm, J Dickinson (O-163) demonstrated their ability to accurately identify the correct ICSI injection site on the oocyte, possibly moving us one step closer to automation of the procedure.
A Mokhtare (O-165) brought us another step closer, reintroducing Sakkas’s microfluidics vision. They developed a semi-automated oocyte denudation microfluidic chip and using murine cumulus-oocytecomplexes demonstrated less shear stress imposed on the oocyte with comparable denudation efficiency, postICSI survival, fertilization, and blastocyst formation rates.
FINAL WORD | SARS-CoV-2
No-one could ignore the impact that SARS-CoV-2 has had on the field of assisted reproduction this year. For one, it was the reason we were sitting at home in front of our computers rather than taking in the sights of Copenhagen and meeting with friends and colleagues we only get the opportunity to see at such events. The impact of the virus was raised both in the opening ceremony and in the main programme, as we continue to adapt our everyday practices and learn of further implications.
Concerning SARS-CoV-2, the importance of the ability to test, track and trace, to prevent the spread of the virus among our staff was highlighted, to enable clinics to continue operating.
W Essahib (O-319) and her team confirmed the presence of ACE2 receptors in human oocytes and on all lineages of pre-and peri-implantation blastocysts, potentially indicating the earliest stages of embryonic development are vulnerable to the SARS-CoV-2 viral infection, mandating monitoring of pregnancies and neonatal outcomes during the pandemic.
Further to this, G Bahadur’s (O-316) retrospective analysis of studies following COVID-positive women and their babies showed evidence for alterations in placental histology, miscarriage, presence of the virus in placental tissue, and maternalfetal vertical transmission during pregnancy.
The take-home message is that the virus will continue to impact our practice, and possibly even in ways we are not yet aware of, as we continue to learn more.
 Colleen Lynch MSc
Colleen Lynch MSc
Training and Scientific Manager
Colleen started working in the embryology field in 2004, following the completion of her undergraduate studies in Genetics and post graduate studies in Medical Genetics. She is currently working towards her PhD in Preimplantation Genetics under Professor Darren Griffin at the University of Kent. Having worked as a both a Clinical Embryologist and Genesis Genetics PGT Laboratory Manager, she joined CooperGenomics™ full time in 2017. Colleen joined CooperSurgical Fertility and Genomics Solutions as Training and Scientific Manager in 2018, with specific focus on training in trophectoderm biopsy and supporting lab teams on its introduction.
Colleen has lectured on PGT at many international conferences and has co-authored several scientific papers and book chapters. Additionally, she has extensive knowledge of ISO:15189 standards, works with UKAS as a technical expert, and is an honorary lecturer at Manchester Metropolitan University on the Clinical Science MSc for the Scientist Training Programme (STP) in Reproductive Science.

 My Clinic is in the United States
My Clinic is in the United States My Clinic is in Canada
My Clinic is in Canada